You can purchase our Software and Reagent Products either on-line (PayPal or Credit Card) or by Purchase Order. For Purchase Orders, please request a quote through our Contact Page.
1. Subscription: MutationForecaster genome interpretation system

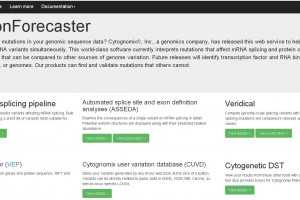
Automated Splice Site and Exon Definition Analysis Server (ASSEDA)
ASSEDA has been incorporated into Cytognomix’s MutationForecaster system. ASSEDA is described in Prediction of Mutant mRNA Splice Isoforms by Information Theory-Based Exon Definition by Mucaki et al. 2013.
Shannon pipeline for mRNA splicing and transcription factor binding site mutation analysis (SP)
SP has been incorporated into Cytognomix’s MutationForecaster system. SP for splicing mutations is described in Interpretation, Stratification and Evidence for Sequence Variants Affecting mRNA Splicing in Complete Human Genome Sequences (Shirley et al. 2013), for transcription factor binding sites in Mucaki et al. 2016, Caminsky et al. 2016, and Lu et al. 2016.
Veridical, for verification of predicted mRNA splicing mutations
Veridical sets the standard for evidence-based, statistical validation of mutation predictions from the Shannon pipeline – or other sources. Veridical has been incorporated into the MutationForecaster system. Veridical is described in Validation of predicted mRNA splicing mutations using high-throughput transcriptome data by Viner et al. 2014.
Variant effect predictor (VEP)
Predict of the effect of variants on coding regions of genes and protein sequence. SIFT and PolyPhen scores are also generated.
Cytognomix user variation database (CUVD)
Store results generated by any of our web tools at the click of a button for variants present in your sample. Variants can be directly related to public data in HGNC, NCBI, EBI, ClinVar, as well as locus specific databases. Based upon the Leiden Open Variation Database.
Cytogenetic Visual Analytic Decision Support Tool (CVDST)
Link literature evidence for gene mutations detected with VEP and SP with this tool. View your results from these other tools with our built-in genome browser. This tool also provides tracks for Cytognomix FISH probes and capture reagents.
After registration with MutationForecaster, subscriptions can be activated through the Account menu on the system.
2. Reagents: BRCA complete gene sequence capture enrichment for next generation sequencing
Capture the complete sequences of 21 genes that are mutated in women at high risk for developing breast or ovarian cancer. Derived with Cytognomix’s patented ab initio probe technology. Performance of these products for capturing exonic, intronic, upstream and downstream sequences has been established in multiplexed NGS. Contact us at the address below for a quote. Data generated with these reagents are analyzed with Cytognomix’s companion software products.
3. Automated Dicentric Chromosome Identifier and Dose Estimator (ADCI).
Software that accelerates the cytogenetic biodosimetry of exposures to various qualities of radiation. The software fully automates the dicentric chromosome (DC) analysis from blood and integrates dose assessment. It has been developed for multiple individuals exposed in mass casualty or moderate scale industrial radiation events, but also can be effectively used to quickly analyze accidents involving one or a few individuals. ADCI offers speed, accuracy, and scalability that solves the challenges that users face to meet the productivity requirements for a mass casualty event. It enables greater standardization between laboratories, while still allowing different labs to customize their own calibration curves for determining unknown radiation exposures, which addresses differences in chromosome preparation methods and radiation calibration sources between labs.
4. Single copy genomic probes: FISH and genomic microarrays
Single copy FISH probes and genomic microarrays may be purchased upon request.
Contact us: info@scprobe.info