Peter Rogan 2015 May 04 6:14 p.m.
The Logic and Formulation of Exon Definition for Splice and Splicing Regulatory Sites with Negative Information Content. PK Rogan, EJ Mucaki
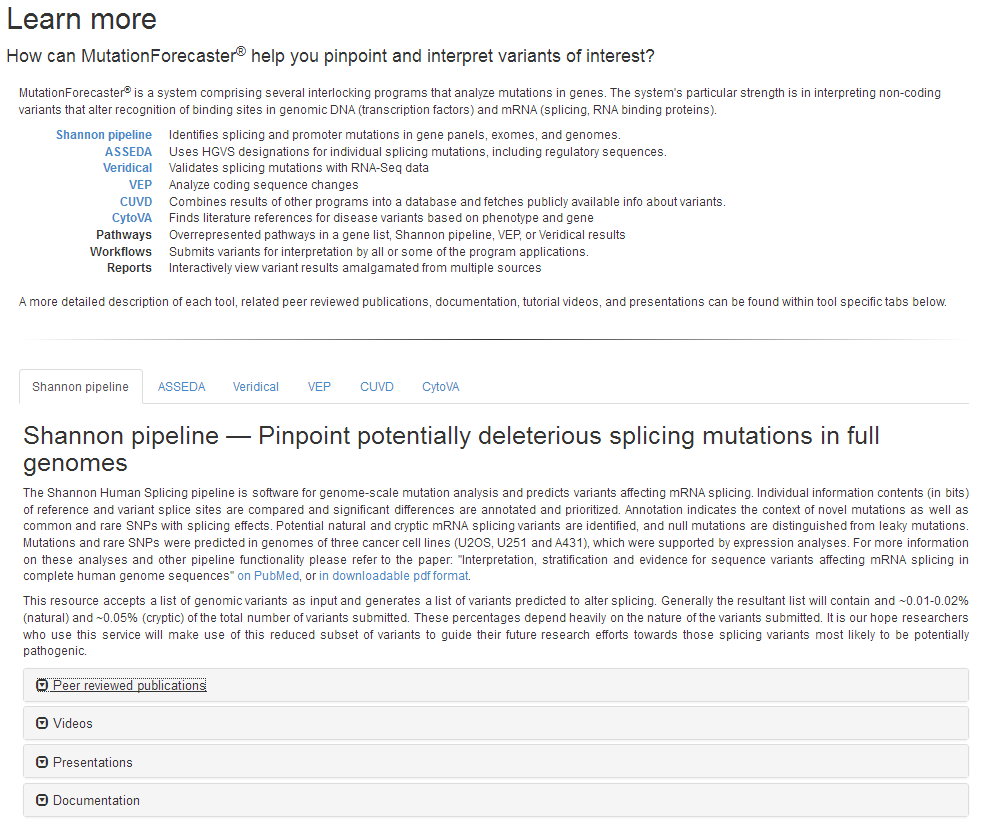
Update on: Mucaki EJ, 2013 and the Automated Splice Site and Exon Definition Analysis server (ASSEDA).
In Mucaki EJ, 2013, we described a method of predicting the overall strength of an exon by calculating its total information content (Ri,total) from the sum of the Ri values of its donor and acceptor splice sites, adjusted for their gap surprisal (the self-information of the distance between the two sites). Differences between ΔRi,total values are predictive of the relative abundance of these exons in distinct processed mRNAs.
Splice sites altered by mutations that prevent stable interaction with splicesomes are said to be abolished. Information theory predicts abolition of binding below their minimum binding affinity, Ri,minimum, which is empirically derived. This value is slightly above zero bits, the theoretical minimum for binding at equilibrium (ΔG = 0; Schneider TD, 1997). Sites with Ri < 0 are not bound, forming stable interactions would be endergonic (ΔG > 0). This raises the question, when predicting the change in exon strength (ΔRi,total) due to a mutation that inactivates binding, whether mutant sites with varying degrees of negative information content are energetically distinguishable from one another.
The computation of Ri,total contains the sum of the the Ri values of component binding sites, irrespective of their initial or final strengths. Thus, a mutated site with Ri << 0 would result in greater ΔRi,total compared to a site with Ri ~ 0. To assess whether the degree of unfavorable binding should be applied to the exon definition calculation, or if values below 0 bits should be computed similarly to a binding site at equilibrium (Ri ~ 0), we reevaluated experimentally validated natural and regulatory splicing mutations in our paper with both approaches. Ri,total was calculated for 10 variants from Supplementary Table 2, both including and excluding the negative information (ie. Ri < 0 vs. Ri = 0) of inactivated splice sites. Mutation #2 of Supplementary Table 2 [ADA:g.43249658G>A] abolishes a natural donor site, from 8.8 to -9.9 bits. In applying the full decrease in strength (ΔRi,total: -18.7 bits), the natural exon strength decreases from 21.0 to 2.3 bits. When the negative information content is set to zero bits, the change is significantly smaller (21.0 -> 12.2 bits; ΔRi,total = -8.8 bits). When a weak natural splice site is abolished, the difference as expressed as ΔRi,total can be quite small (Mutation #9; -14.8 vs -3.1 bits). In the case of Mutation #38, the reduction in ΔRi,total leads to a partially discordant prediction where the abolished natural exon is weaker than the experimentally confirmed activated cryptic exon. Results for this mutation were concordant with the published version when the negative bit value of the mutated natural site was included in the calculation.
The impact of mutations in splicing regulatory (SR) factors can also be predicted on ASSEDA, where the Ri of the SR binding site is added to the R_i,total, as well as a secondary gap surprisal value for the particular SR protein. These sites can also be abolished. But when a SR protein binding site is no longer active, should the SR gap surprisal still be applied, or is the SR gap surprisal no longer applicable?
We tested mutations from Mucaki EJ, 2013 (Supplementary Table 4), which abolish the splicing enhancer SF2/ASF with and without the SR protein gap surprisal when Ri of the SR site is < 0 bits. The removal of the gap surprisal term for Mutation #2 of Supplementary Table 4 leads to a discordant prediction, where the ΔRi is less than the SR gap surprisal at that distance and therefore the ΔRi,total is positive. As experimental evidence shows an increase in skipping, it is a discordant prediction. Therefore, the gap surprisal is still applied in the computation of both initial and final Ri,total values when the SR protein of interest is abolished as the site is naturally present and therefore expected for binding. Conversely, when we apply the gap surprisal to the initial Ri,total for a splicing factor that is being created, we are essentially applying a penalty for a site that does not normally exist. Therefore, we no longer apply the SR gap surprisal value to the initial Ri,total in these cases.
The revised Ri,total values of SR binding site mutations slightly differ from those reported in Mucaki EJ, 2013 (Supplementary Table 4). This is because the gap surprisal distributions were recomputed for the following factors: SF2/ASF, SC35 and SRp40, with updated versions of these models based on CLIP Seq data (Blin K, 2015, Khorshid M, 2011). This resulted in small changes to the distributions for SF2/ASF and SC35, however changes for SRp40 were significant, and now more closely resembles the other gap surprisal functions. The updated graphs of distance vs. gap surprisal are available at: http://splice.uwo.ca/gapsurprisals.html. While this should not significantly affect ΔRi,total values, it may affect the initial and final Ri,total values.
-
This article was mentioned in a comment by Peter Rogan2018 Jan 12 2:39 p.m.
-
This article was mentioned in a comment by Peter Rogan2017 Dec 12 09:53 a.m.
See:Analysis of the effects of rare variants on splicing identifies alterations in GABAA receptor genes in autism spectrum disorder individuals.[Eur J Hum Genet. 2013.]
-
This article was mentioned in a comment by Peter Rogan2017 Dec 07 5:24 p.m.
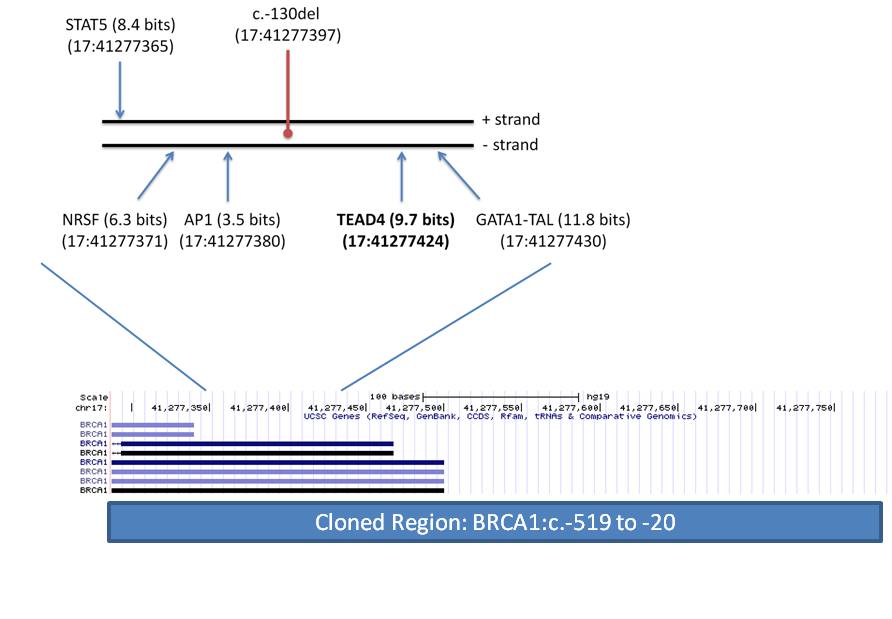
See:Characterization of a novel germline BRCA1 splice variant, c.5332+4delA. [Breast Cancer Res Treat. 2017.]
-
This article was mentioned in a comment by Peter Rogan2017 Oct 01 8:57 p.m.
See:Rules and tools to predict the splicing effects of exonic and intronic mutations. [Wiley Interdiscip Rev RNA. 2018.]