Author Archives: Peter Rogan
December 14. New paper published about regulation of gene transcription by information theory and machine learning
Lu R and Rogan PK. Transcription factor binding site clusters identify target genes with similar tissue-wide expression and buffer against mutations [version 1 ]. F1000Research 2018, 7:1933 (DOI: 10.12688/f1000research.17363.1): https://f1000research.com/articles/7-1933/v1
December 7, 2018. New article describing database of validated mRNA splicing mutations published
Shirley BC, Mucaki EJ and Rogan PK. Pan-cancer repository of validated natural and cryptic mRNA splicing mutations [version 1; referees: awaiting peer review]. F1000Research 2018, 7:1908 (https://doi.org/10.12688/f1000research.17204.1
Website to perform database search: https://validsplicemut.cytognomix.com/
November 28, 2018. New automated cytogenetic biodosimetry article accepted for publication
RADIATION DOSE ESTIMATION BY COMPLETELY AUTOMATED INTERPRETATION OF THE
DICENTRIC CHROMOSOME ASSAY
Li, Yanxin; Shirley, Ben; Wilkins, Ruth; Norton, Farrah; Knoll, Joan; Rogan,Peter
Radiation Protection Dosimetry, in press
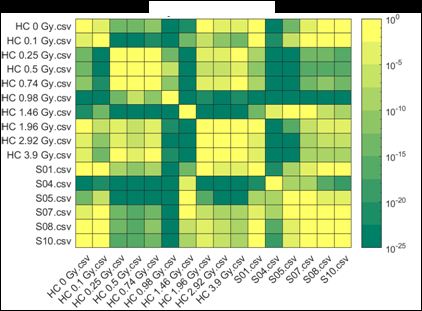
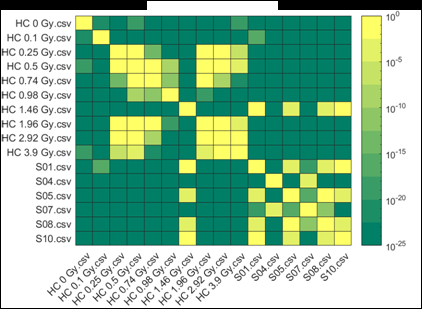
Figure 2 (from the article): Representative heat maps of chromosome object count distributions for Health Canada calibration (HC ##Gy.csv) and exercise (S##.csv) samples. Show: legend indicates significance of pairwise comparisons of object counts in (bottom panel) unselected vs. (top panel) Model B_C filter (5) selected images from each sample. Values close to 10^0 are not significant, whereas values such as 10^-20 are significant differences between samples. The legend displays probability thresholds of Wilcoxon signed rank tests of chromosome object counts between each pair of samples from the HC laboratory. Similar results were obtained for other chromosomal features.
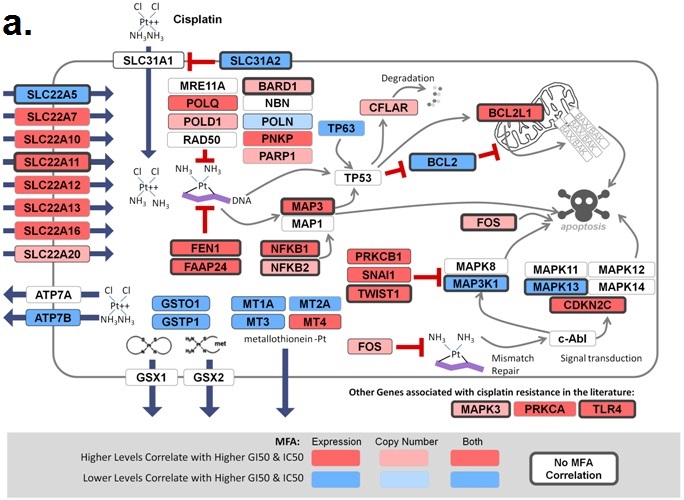
November 6, 2018. Article on prediction of platin chemotherapy response accepted for publication
Mucaki EJ, Zhao J, Lizotte D, Rogan PK. Predicting response to platin chemotherapy agents with biochemically-inspired machine learning. Signal Transduction and Targeted Therapy, in press.
A preprint of this article has been released on the BioRxiv server: https://doi.org/10.1101/231712
October 19, 2018. Presentations at the 7th Northeast Regional Chromosome Pairing Conference
CytoGnomix and the University of Western Ontario presented several papers about differential accessibility of single copy FISH probes to metaphase chromosomes at the Chromosome Pairing conference, held at Laurentian University, Sudbury, Ontario, Canada. We are grateful to Prof. Thomas Merritt for inviting us to participate in this exciting conference. The following talks were presented:
Seana Hill (Knoll and Rogan labs): Defining differentially accessible domains within human
metaphase homologues.
Jerry Wang (Knoll and Rogan labs): Expanding the catalog of differentially compacted loci
on homologous chromosomes in the human genome.
Peter Rogan (Rogan lab): A proposed mechanism to establish differential allelic
accessibility to homologous metaphase chromosomes.
Sept. 11, 2018. New publication about promoter gene variants in BRCA1 and BRCA2 that alter gene expression
In a collaboration with the ENIGMA Consortium members, we have used bioinformatic and functional genomic analysis to identify gene variants that affect the expression of the BRCA1 and BRCA2 genes. The article is:
Burke LJ, Sevcik J, Gambino G1, Tudini E, Mucaki EJ, Shirley BC, Whiley P, Parsons M, DeLeeneer K, Gutiérrez-Enríquez S, Pena MS, Caputo S, Santana dos Santos E, Soukupova J, Janatova M, Zemankova P, Lhotova K, Stolarova L, Borecka M, ENIGMA consortium, Edwards S, Blok R, Hansen TvO, Diez O, Vega A, Claes K, Rouleau E, , Radice P, Peterlongo P, Rogan PK, Caligo M, Spurdle AB and Brown MA. BRCA1 and BRCA2 5’ non-coding region variants identified in breast cancer patients alter promoter activity and protein binding, Human Mutation.
Link to full text of accepted paper: http://www.cytognomix.com/?attachment_id=4541
Permanent link: https://doi.org/10.1002/humu.23652
Aug. 13, 2018. Differential accessibility to homologous chromosomal loci confirmed by international consortium
A large international consortium based at Harvard University has demonstrated parental homolog-specific differences in chromatin accessibility on human chromosome 19:
Nir et al. BioRxiv doi: 10.1101/374058 (Walking along chromosomes with super-resolution imaging, contact maps, and integrative modeling)
This work reproduces previous reports previously published by CytoGnomix scientists using our patented scFISH™ probes:
Khan et al. Molecular Cytogenetics 2014 7:70 (Localized, non-random differences in chromatin accessibility between homologous metaphase chromosomes)
Khan et al. Molecular Cytogenetics 2015 8:65 (Reversing chromatin accessibility differences that distinguish homologous mitotic metaphase chromosomes)
June 13, 2018. Article in Fast Forward
The 2018 Impact report from the Southern Ontario Smart Computing for Innovation Platform (SOSCIP), which supports the development of a supercomputer version of the Automated Dicentric Chromosome and Dose Estimation (ADCI) system in IBM Blue Gene/Q, contains an article about our project:
https://www.soscip.org/wp-content/uploads/2017/08/soscip_impactreport2018_pages.pdf
June 7, 2018. Presentations at upcoming international conferences
Population scale biodosimetry with the Automated Dicentric Chromosome Identifier and Dose Estimator (ADCI) software system. (Platform) Rogan PK, Ali, S, Li Y, Shirley B, Wilkins R, Flegal F, Cooke R, Peerlaproulx T, Waller E, Knoll JHM. EPR Biodose 2018, June 11-15, Munich Germany
Optimization of image selection in Automated Dicentric Chromosome Analysis. Li Y, Shirley B, Wilkins R, Flegal, F, Knoll JHM, Rogan PK. EPR Biodose 2018, June 11-15, Munich Germany.
Predicting exposure to ionizing radiation by biochemically-inspired genomic machine learning. Rogan PK, Zhao JZL, and Mucaki EJ. EPR Biodose 2018, June 11-15, Munich Germany.
Comprehensive prediction of responses to chemotherapies by biochemically-inspired machine learning. (Best Poster session) Rogan PK, Zhao JZL, Mucaki EJ. European Society of Human Genetics 2018, June 16-19, Milan Italy.
May 29, 2018. Change to cytognomix.org URL
As of today, we have transitioned the website cytognomix.org to:
http://academic.cytognomix.com
The site contains our published articles, lectures and presentations about human genetics and molecular biology. All of the legacy content at this site (1980-2007) will be preserved on the new site.
Please update your browser bookmarks to reflect this change.
March 29, 2018. Mutation Forecaster: Key to discovery of mutations in novel ALS gene
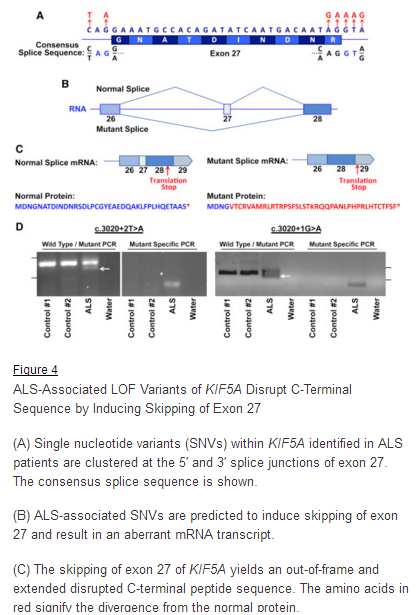
CytoGnomix’s Mutation Forecaster: Key to discovery of mutations in novel ALS gene. Nicolas et al. Genome-wide Analyses Identify KIF5A as a Novel ALS Gene. Neuron 97:1268-1283.e6, 2018
http://www.cell.com/neuron/fulltext/S0896-6273(18)30148-X
From our subscriber, Dr. John Landers (U. Mass. School of Medicine):
“We used the application ASSEDA (Automated Splice Site and Exon Definition Analyses) to predict any mutant mRNA splice isoforms resulting from these variants (Mucaki et al. 2013). This algorithm was chosen as it is known to have high performance in splice prediction (Caminsky et al., 2014). ASSEDA predicted a complete skipping of exon 27 for all variants, yielding a transcript with a frameshift at coding amino acid 998, the deletion of the normal C-terminal 34 amino acids of the cargo-binding domain, and the extension of an aberrant 39 amino acids to the C terminus (Table 3; Figures 4B and 4C). The presence of transcripts with skipped exon 27 was demonstrated by performing…”
April 9, 2018. Upcoming release of Automated Dicentric Chromosome Identifier and Dose Estimator (ADCI)
Request a quote.
March 23, 2018. New preprint about target gene regulation by transcription factors
Clustered, information-dense transcription factor binding sites identify genes with similar tissue-wide expression profiles. BioRxiv, 2018.
March 13, 2018. Oral presentation on chemotherapy response in Best Poster session at ESHG 2018
On behalf of the Scientific Programme Committee of the European Conference of Human Genetics 2018 taking place in Milan, Italy from June 16 to June 19, 2018, we are pleased to inform you that the abstract entitled:
‘Comprehensive prediction of responses to chemotherapies by biochemically-inspired machine learning’
(Control No. 2018-A-2095-ESHG)
was among the best scored papers accepted for a poster presentation. Best Poster session takes place on Sunday, June 17, 2018 13:00 hrs, and consists of a 3 minute presentation followed by discussion at your electronic poster.
February 28, 2018. New publication on radiation biodosimetry based upon machine learning
We have published a new approach to devise gene signatures to detect radiation exposure (human, murine), and to quantify levels of exposure (murine):
Zhao JZL, Mucaki EJ and Rogan PK. Predicting ionizing radiation exposure using biochemically-inspired genomic machine learning. F1000Research 2018, 7:233 (doi: 10.12688/f1000research.14048.1)
February 19, 2018. Article on genomic signature of radiation exposure
Manuscript describing accurate genomic signatures of radiation exposure will be published shortly by F1000Research.
Jonathan ZL Zhao, Eliseos J Mucaki, Peter K Rogan. Predicting Exposure to Ionizing Radiation by Biochemically-Inspired Genomic Machine Learning, F1000Research, in press.
Abstract:
Background: Gene signatures derived from transcriptomic data using machine learning methods have shown promise for biodosimetry testing. These signatures may not be sufficiently robust for large scale testing, as their performance has not been adequately validated on external, independent datasets. The present study develops human and murine signatures with biochemically-inspired machine learning that are strictly validated using k-fold and traditional approaches.
Methods: Gene Expression Omnibus (GEO) datasets of exposed human and murine lymphocytes were preprocessed via nearest neighbor imputation and expression of genes implicated in the literature to be responsive to radiation exposure (n=998) were then ranked by Minimum Redundancy Maximum Relevance (mRMR). Optimal signatures were derived by backward, complete, and forward sequential feature selection using Support Vector Machines (SVM), and validated using k-fold or traditional validation on independent datasets.
Results: The best human signatures we derived exhibit k-fold validation accuracies of up to 98% (DDB2, PRKDC, TPP2, PTPRE, and GADD45A) when validated over 209 samples and traditional validation accuracies of up to 92% (DDB2, CD8A, TALDO1, PCNA, EIF4G2, LCN2, CDKN1A, PRKCH, ENO1, and PPM1D) when validated over 85 samples. Some human signatures are specific enough to differentiate between chemotherapy and radiotherapy. Certain multi-class murine signatures have sufficient granularity in dose estimation to inform eligibility for cytokine therapy (assuming these signatures could be translated to humans). We compiled a list of the most frequently appearing genes in the top 20 human and mouse signatures. More frequently appearing genes among an ensemble of signatures may indicate greater impact of these genes on the performance within individual signatures. Several genes in the signatures we derived are present in previously proposed signatures.
Conclusions: Gene signatures for ionizing radiation exposure derived by machine learning have low error rates in externally validated, independent datasets, and exhibit high specificity and granularity for dose estimation.
February 7, 2018. Accepted presentations at EPR Biodose (Munich, June, 2018)
Ali, S, Li Y, Shirley B, Wilkins R, Flegal F, Rogan PK, Knoll JHM. Population scale biodosimetry with the Automated Dicentric Chromosome Identifier and Dose Estimator (ADCI) software system. [Platform]
Rogan PK, Zhao JZL, and Mucaki EJ. Predicting exposure to ionizing radiation by biochemically-inspired genomic machine learning.[Poster]
Li Y, Shirley B, Wilkins R, Flegal, F, Knoll JHM, Rogan PK. Optimization of image selection in Automated Dicentric Chromosome Analysis. [Poster]