Mutation interpretation software.
Our MutationForecaster suite is designed to analyze individual variants, gene panels, complete exomes or genomes. One component is our Shannon pipeline for splicing mutation analysis, the only product on the market capable of comprehensive prediction of mRNA splicing mutations in exomes or complete genomes. This product finds mutations that no other software, public domain or commercial, detects. It is based on a proven technology backed up by more than 400 peer-reviewed publications, and thousands of users. Aside from the online cloud application, it is available as an Application on Illumina’s BaseSpace system. Veridical is another genome-scale product for the automatic validation of DNA sequencing variants that alter mRNA splicing. It is companion software to the article “Validation of predicted mRNA splicing mutations using high-throughput transcriptome data”, Viner et al. published in F1000research. Veridical performs statistically valid comparisons of the normalized read counts of abnormal RNA species in mutant versus non-mutant tissues using RNASeq data. Veridical uses output from the Shannon human splicing pipeline as input. The Automated Splice Site and Exon Definition Analysis server predicts the effects of sequence changes that alter mRNA splicing in human diseases. We designed the system to evaluate changes in splice site strength based on information theory-based models of donor and acceptor splice sites. It has been updated with the latest genomic coordinates (HG19), the newest mRNA tables (UCSC), known SNPs (dbSNP135), and updated donor and acceptor weight matrices. ASSEDA predicts potential normal and mutant isoform structures and their relative abundance.
Solution capture hybridization reagents.
Oligonucleotide pools covering unique sequences present in promoter, intragenic, and downstream regions have been designed for a series of genes. The first product has been validated for 7 hereditary breast and ovarian cancer genes. These reagents enable targeted enrichment of specific genes in any next generation, high throughput sequencing platform.The coverage using ab initio technology extends beyond products available from others, since Cytognomix covers both repeat-masked and divergent repetitive sequences with properties of unique sequences. This means that high multiplicity coverage is available for regions that were previously avoided – regions potentially containing mutations that would have been missed by other commercially available capture libraries.
Radiation biodosimetry software.
The Automated Dicentric Chromosome Identifier and Dose Estimator (ADCI) completely automates the interpretation of cytogenetic biodosimetry data. The performance of the system fulfills International Atomic Energy Agency criteria for triage biodosimetry. Both workstation and high performance computing versions can accurately estimate exposures using image data obtained from any commercial metaphase system on the market, without user curation. See our biodosimetry product page for more details. You can also test our algorithm for discriminating dicentric from monocentric chromosomes.
Laboratory-validated SCFISH probes.
The applicability and precision of the SC-FISH has been established for over twenty different chromosomal disorders, covered by over seventy different probes, and all currently available. This set includes probes for congenital syndromes and leukemias, for conditions such as Monosomy 1p36, Wolf-Hirschorn, Cri-du-Chat, Chronic Myelogenous Leukemia (CML), Acute Lymphoblastic Leukemia (ALL), Acute Myeloid Leukemia (AML), Prader-Willi & Angelman, Smith-Magenis Syndromes, and many others. A complete set of subtelomeric SCFISH probes has also been validated for detection of chromosome imbalances close to the termini of individual chromosomes.
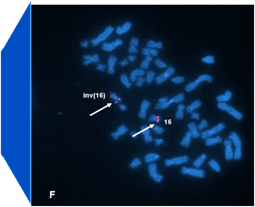
Diagnostic results from a patient with acute myelogenous leukemia
The Cytognomix SC Genome Browser.
The Company’s SC Genome Browser contains design information on over 220,000 ab initio single-copy intervals for which SC FISH probes have been pre-designed or are being used for microarray content development. The probes can be selected by chromosomal location (band or genomic coordinate), by gene locus, and by disease or phenotype. Each probe is unique, and is representative of a single–copy interval, as assigned by the Company’s proprietary ab initio processes. Please visit the browser website or request an account for access to these products.
Microarray product designs for array comparative genomic hybridization.
Array CGH is used to identify chromosome abnormalities by genome-wide assessment of sequence copy number. Fundamental questions remain about data quality, evident by inter-platform comparisons monitored by the US FDA. Coefficients of variation remain high between platforms. We have shown that noise in genomic hybridization is related to contaminating SC sequences present in repeat-enriched (C0t-1 DNA) used to block repetitive sequences, and repetitive sequences in local context to single copy probes on those arrays. Systematic noise in microarray data can be reduced with ab initio SC designs, eliminate the requirement for C0t-1 DNA and obviate its effects.
Repeat-masked probes.
Cytognomix’s founders previously invented and carried out studies based on repeat-masked approaches* for SCFISH probe design. In most instances, these probes perform as expected, however, we have found that only ab initio technology can assure true genome-wide SC designs.
