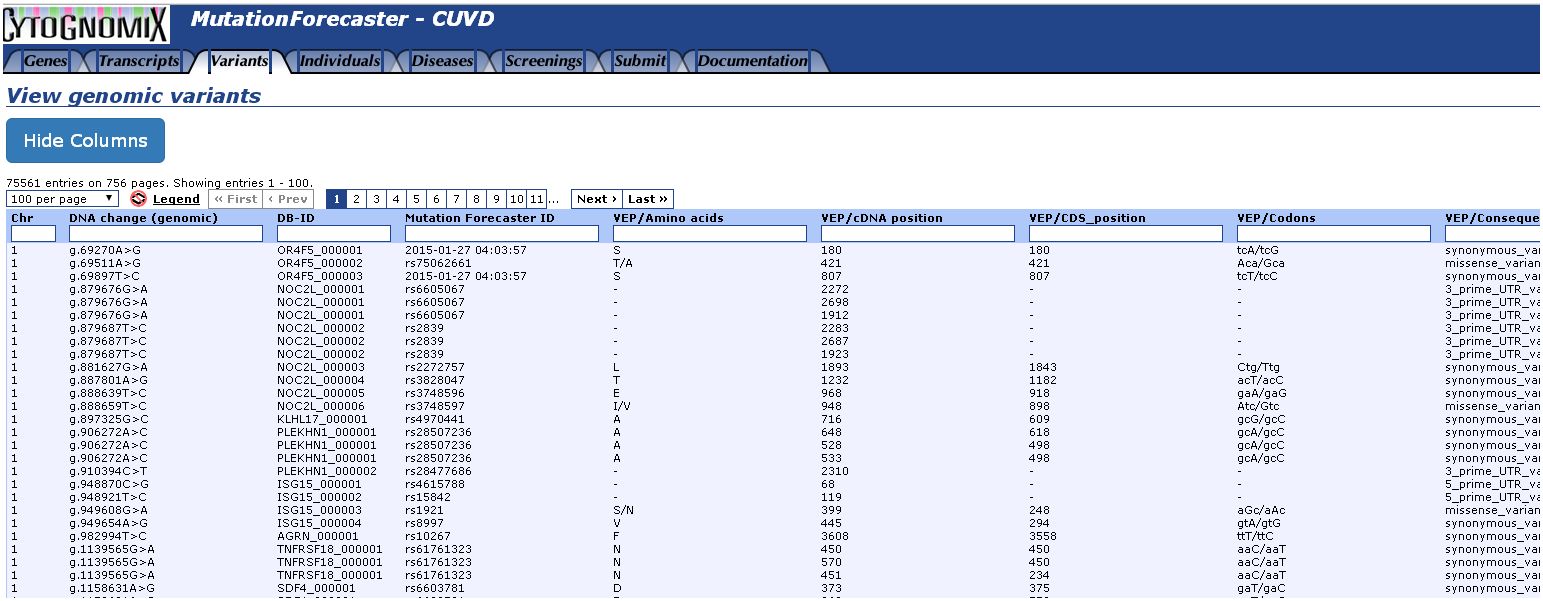
Developed from the widely-used Leiden Open Variation Database, CUVD stores and manages the results of variant analysis by Automated splice site and exon definition analysis (ASSEDA), Shannon pipeline, Variant effect predictor (VEP), and Veridical. It it editable and is also capable of performing built-in realtime searches of other locus-specific databases, Clinvar, dbSNP, and the Exome Variant Server. All samples can be searched for the same variant, or variants can be retrieved for individual samples.
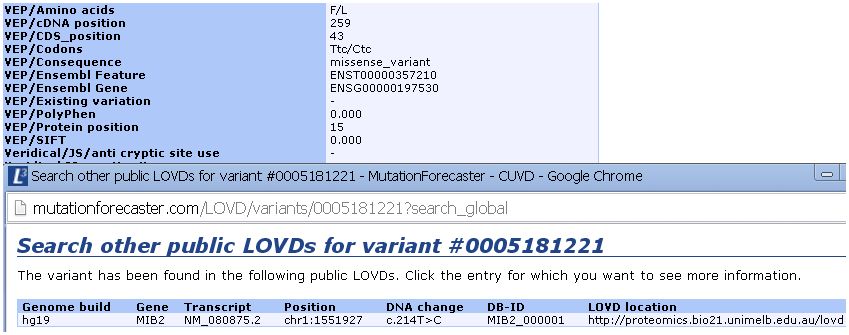
Below, we have imported results from VEP from a breast tumor exome and searched other databases for a missence change in the MIB2 gene: