Peter Rogan, President of Cytognomix, accepted a Certificate of Appreciation from Hon. Sylvia Jones, Deputy Premier and Minister of Health of the Province of Ontario for our work on community COVID-19 outbreaks and their risk factors. This work was carried out in response to the Ministry’s Challenge Questions initiative to researchers, clinical professionals, hospitals and companies addressing how to approach this disease. Our work contributed to answering the challenge questions: Can we better predict the trajectory of COVID-19 across multiple pandemic waves? What are the drivers of future pandemic waves?
In the letter that accompanied the certificate, the Minister wrote:
Thanks to your research, we are able to understand the importance of early identification of probable COVID-19 hotspots and the available mitigation strategies to limit rapid spread of COVID-19. Having contributions such as yours allows us to get the right information to the right people at the right time to support the COVID-19 pandemic response.
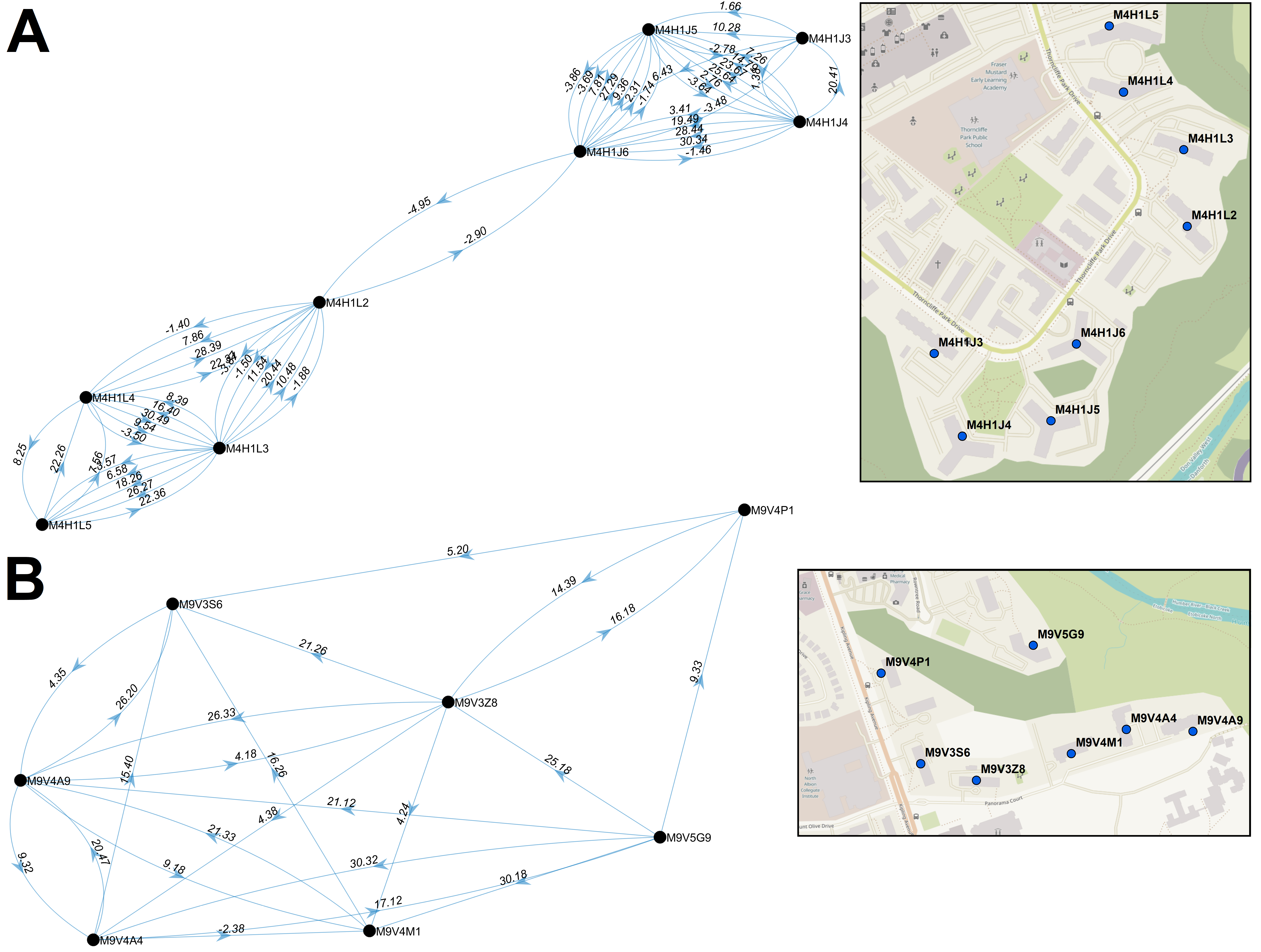
Our geostatistical epidemiology analyses of incidence and locations of infected individuals during Waves 1-3 provided the Ministry with key insights to support Ontarians during the COVID-19 pandemic.